Recent advances in light microscopy are raising the bar on image resolution. Increasingly, many journal reviewers are requesting high-resolution images to accompany their published studies, says Craig Brooks, Ph.D., a cell biologist and microscopy expert at Vanderbilt University Medical Center.
Last year, collaborators at Harvard University reached out to Brooks to acquire higher resolution imaging of kidney tissue. This led him to try an unconventional microscopy technique to image organelles inside ischemically injured proximal tubule cells. After the images published, the editors of Methods in Cell Biology asked Brooks to outline his technique in a special edition on kidney disease.
“This technique wasn’t originally designed for tissue. It’s mainly set up for cultured cells, that are narrow, flat and growing on glass or plastic,” Brooks said. “But we’ve tried it in a variety of tissues, and it’s worked.”
How Microscopy Works
Brooks explained the technique, called structured illuminated microscopy (SIM), is a kind of go-between electron microscopy and light microscopy. It requires no special dyes or buffers.
“Structured illumination microscopy instead relies on shooting a laser through a series of slits to generate a pattern on the sample. From that pattern and the signal it creates, we can decipher a higher resolution than is normally possible by light microscopy,” Brooks said.
“Traditionally, light microscopy cannot separate objects less than 200 nm apart, but with SIM we can double that resolution to 100 nm.”
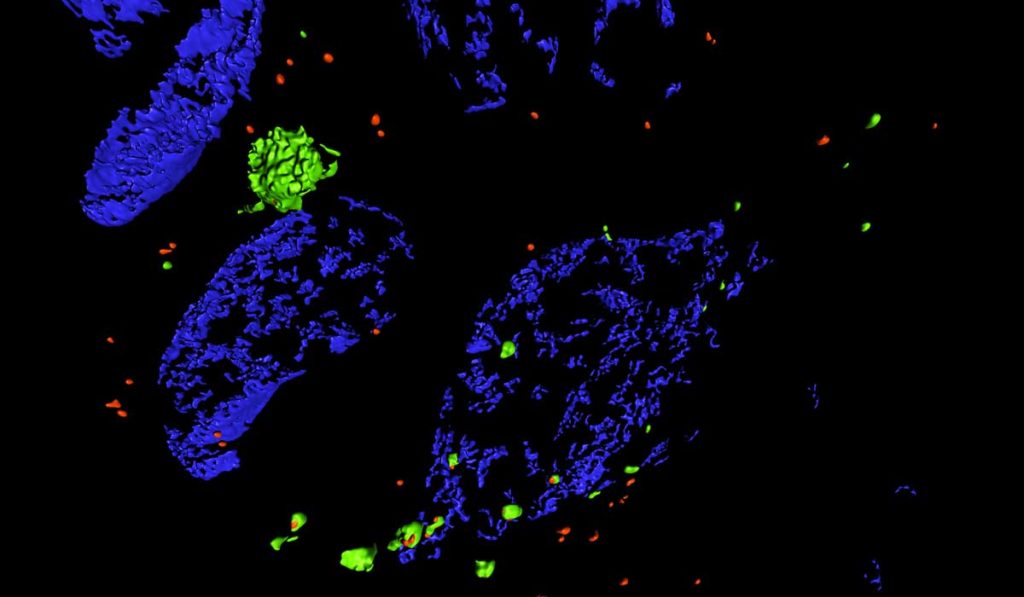
Researchers can use SIM to view tissue sections that have been labeled with fluorescent antibodies. “It allows us to look at organelles that we can usually only see in isolated cells, or a single layer of cells in culture. SIM lets us see the same level of detail but in tissue ,” Brooks said. “Traditionally, light microscopy cannot separate objects less than 200 nm apart, but with SIM we can double that resolution to 100 nm.”
Tracking Autophagy
SIM holds particular promise for not only viewing subcellular organelles, but for monitoring disease-related changes. The new methods paper demonstrates SIM’s ability to measure autophagy dynamics – the cellular “self-eating” mechanism for breaking down faulty intracellular components.
Autophagy plays a central role in cellular stress response. Its dysfunction is associated with myriad pathologies including cancer, aging, cardiovascular disease, liver disease, and of course, kidney disease. A major challenge in studying autophagy has been in reaching consensus on the right way to measure it. Static assays miss autophagic turnover, and it can be hard to differentiate between early autophagosomes and late autolysosomes – particularly when the two fuse inside cells.
The paper helps bridge the gap between assays by showing it is possible to monitor autophagic flux in kidney tissue using SIM. By combining SIM with pH-sensitive fluorophores, Brooks was able to monitor the stages of autophagosome formation.
“We were able to see not just how many autophagosomes there were, but what stages they were in.”
“We were able to see not just how many autophagosomes there were, but what stages they were in,” Brooks said. “With SIM we could tell which had just formed, which were in the lysosome, and which were in the gradient between.”
The paper also includes side-by-side images showing how autophagosomes increased in size after the researchers induced kidney injury in reporter mice. Importantly, the study revealed injuring one kidney causes physiological changes in the other; SIM showed significant autophagic flux in both kidneys just 24 hours after just one kidney was injured.
Broad Implications
The validated microscopy approach opens the door for a host of applications. Brooks envisions SIM supporting molecular-level diagnostics as new fluorescent markers develop.
Take glomerular disease as an example, Brooks said. “Traditionally, glomerular diseases are diagnosed by a pathologist via electron microscopy. But there’s work ongoing to use SIM, and feed detailed images into a computer algorithm to help diagnose based on the staining pattern. In the future, SIM could provide pathologists with another tool to diagnose various diseases.”