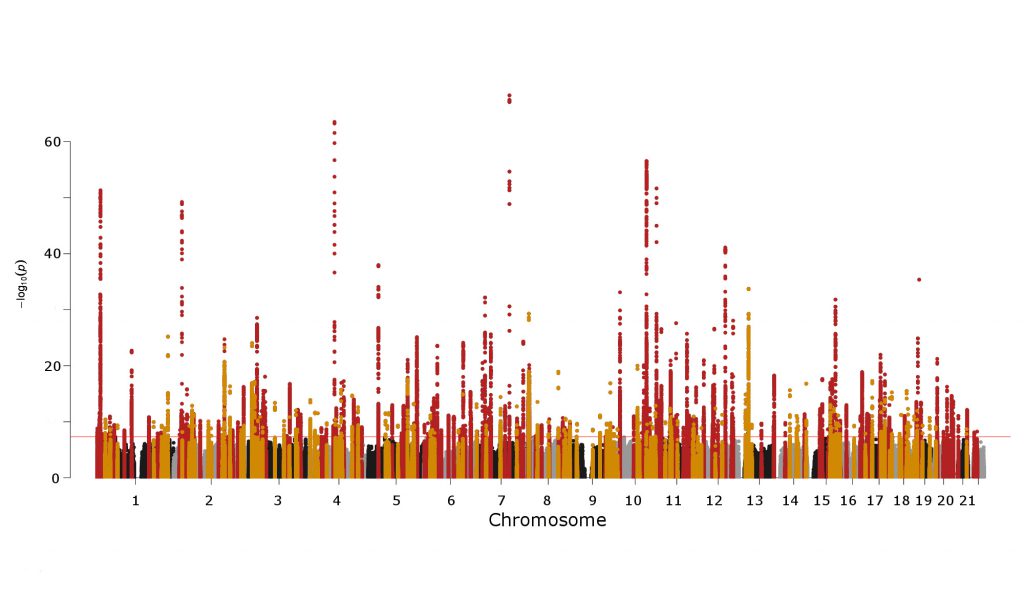
For decades, researchers have been uncovering genetic variations that increase risk of high blood pressure (BP). Hundreds of single nucleotide polymorphisms (SNPs) have been reported. These SNPs can slightly elevate BP risk associated with cardiovascular disease, including coronary artery disease, stroke or kidney disease complications.
While each variant has a tiny effect, some could be modifiable or serve as therapeutic targets. Researchers from the Vanderbilt Genetics Institute have helped develop a comprehensive catalog of SNPs associated with high BP. Published in the journal Nature Genetics, the initiative was the largest of its kind to date, involving an international team of over 100 researchers from around the world.
“Our goal was… to redraw the map of blood pressure genes developed by genetic association studies over the last decade.”
Todd Edwards, Ph.D., associate director of the Vanderbilt Genetics Institute, was a senior author on the study. “Our goal was to use newer methods and collaborations with many groups to redraw the map of blood pressure genes developed by genetic association studies over the last decade,” Edwards said. His co-senior author on the study was Adriana Hung, M.D., a nephrologist and epidemiologist at Vanderbilt.
An Unprecedented-Scale Analysis
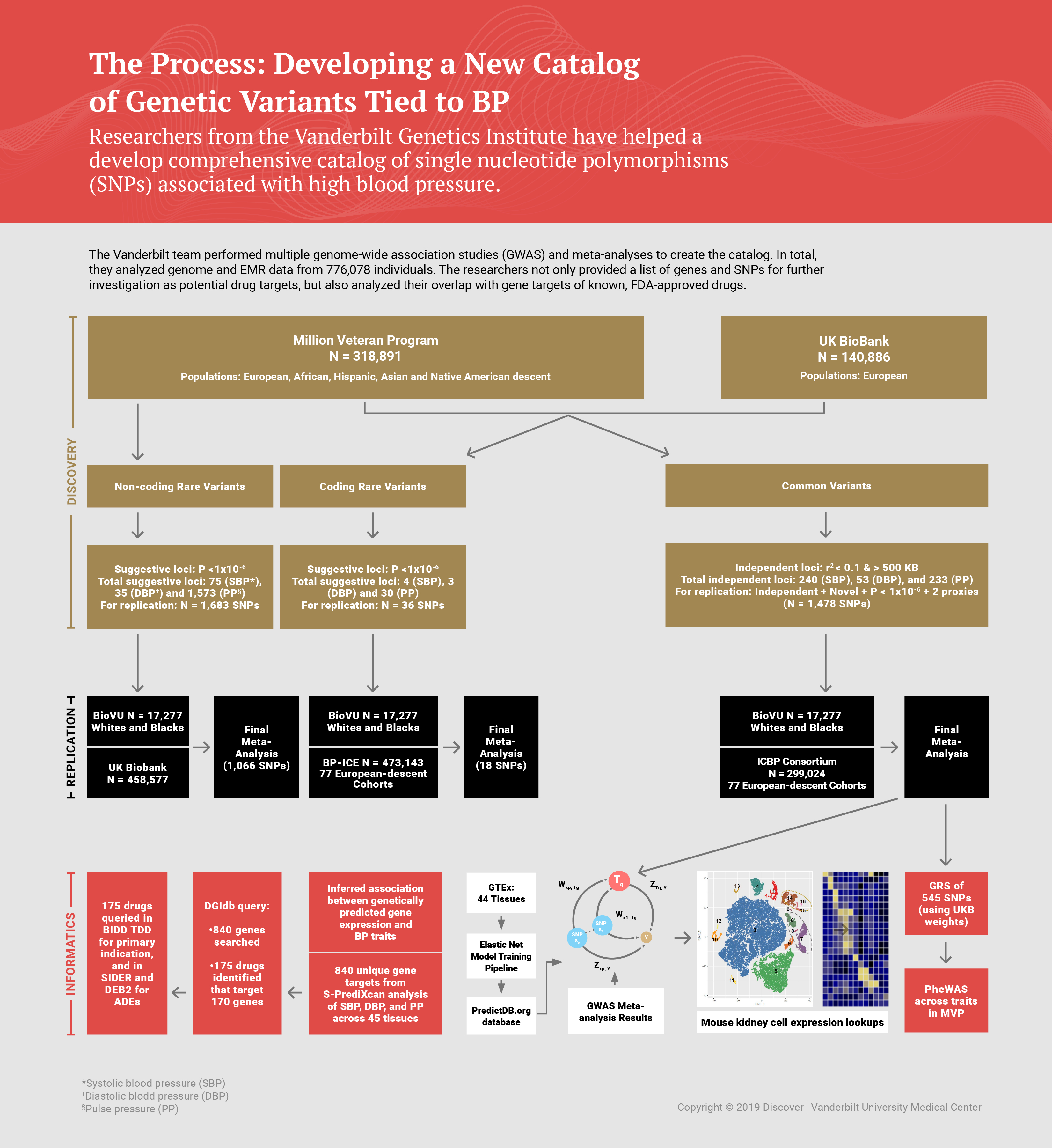
The team performed multiple genomewide association studies (GWAS) and meta-analyses to create the catalog. In total, they analyzed genome and EHR data from 776,078 individuals. They discovered 208 novel, common SNPs and 53 rarer variants associated with high BP.
Analysis began with data from 459,777 participants, 318,891 from the Million Veteran Program (MVP) and 140,886 from the UK Biobank (UKB). The researchers then cross-checked the most significant variant associations in 316,301 participants from the International Consortium for BP and Vanderbilt’s own biobank (BioVU).
One of the major benefits of using the MVP biobank was its diversity. Thirty percent of veterans in the MVP were black, Hispanic, Asian or native American. Biobank diversity helped the researchers find common SNPs that cross ancestries.
From SNP List to Physiology
After finding the SNPs, the team also identified specific tissues where the altered BP genes exert effects.
“We used a method called S-PrediXcan to integrate information from our GWAS with gene-expression information from the Genotype-Tissue Expression (GTEx) project,” said Ayush Giri, Ph.D., assistant professor in the Department of Obstetrics and Gynecology at Vanderbilt, one of four first authors listed on the study. Other first authors from Vanderbilt included postdoctoral fellows Jacklyn Hellwege, Ph.D., and Jacob Keaton, Ph.D.
By comparing genetic variants to regulatory pathways found in healthy tissues, the team predicted how SNPs may affect other aspects of a person’s physiology. Their genotype-tissue expression analyses revealed 840 genes across 45 tissues strongly associated with elevated BP.
The researchers not only provided a list of genes and SNPs for further investigation as potential drug targets, but also analyzed their overlap with gene targets of known, FDA-approved drugs. They found 617 drug-gene interactions that represent ways drugs designed for other purposes might be repurposed as anti-hypertensives with further research.
Translation to Patient Care
The study provides novel insights into potential mechanisms by which genes regulate blood pressure, and is in turn is an opportunity to help patients under treatment for hypertension. It also provides a springboard for future research into conditions related to BP, such as kidney disease, diabetes, inflammation and their secondary complications.
“We are looking at the drugs that target blood pressure genes using electronic health records to explore the potential for repurposing drugs to treat hypertension, as well as identifying new gene targets for drug development,” Edwards said.